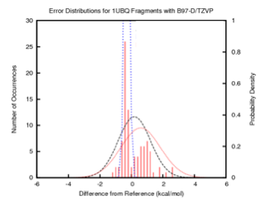
Estimation of Error in Energy Functions
The linked databases were used to develop an estimate of the error of any computational method relative to converged QM calculations. The details of this work are contained in the following manuscripts:
- Merz, K. M., Jr. (2010) Limits of Free Energy Computation for Protein-Ligand Interactions, Journal of Chemical Theory and Computation 6, 1769-1776.
- Faver, J. C., Benson, M. L., He, X., Roberts, B. P., Wang, B., Marshall, M. S., Kennedy, M. R., Sherrill, C. D., and Merz, K. M., Jr. (2011) Formal Estimation of Errors in Computed Absolute Interaction Energies of Protein-Ligand Complexes, Journal of Chemical Theory and Comput. 7, 790-797.
- Faver, J. C., Benson, M. L., He, X., Roberts, B. P., Wang, B., Marshall, M. S., Sherrill, C. D., and Merz, K. M., Jr. (2011) The Energy Computation Paradox and ab initio Protein Folding, Plos One 6, e18868.
- Ucisik, M. N., Dashti, D. S., Faver, J. C., and Merz, K. M., Jr. (2011) Pairwise additivity of energy components in protein-ligand binding: The HIV II protease-Indinavir case, Journal of Chemical Physics 135, 085101.
- John C. Faver, Wei Yang, and Kenneth M. Merz, Jr. (2012) The Effects of Computational Modeling Errors on the Estimation of Statistical Mechanical Variables Journal of Chemical Theory and Computation 8, 10, 3769–3776
Statistical Models for Basis Set Superposition Error
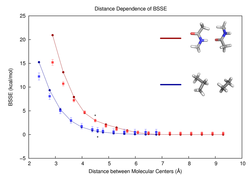
With increased availability of both computing power and clever algorithms, quantum mechanical methods are now being applied to large biomolecular systems. Because such calculations necessarily use incomplete basis sets, QM energies contain basis set incompleteness error. When these methods are applied to large folded systems such as proteins, basis set incompleteness leads to very high magnitudes of intramolecular basis set superposition error(IBSSE). IBSSE is traditionally estimated by fragmenting a supersystem into N small molecular components and performing a counterpoise calculation for each, leading to a total of 2N+1 separate QM calculations. In order to bypass the additional 2N QM calculations, we developed a statistical model trained with molecular fragments found in the Protein Databank. You can download the dataset here. By using this model, one can compute just one QM energy for the supersystem and very quickly get an estimate (with uncertainty) of the IBSSE. There is a web app for fragment-based BSSE estimation available (see drop down menu) along with its source code. A fuller featured version is being developed.
- Faver, J. C., Zheng, Z., and Merz, K. M., Jr. (2011) Model for the fast estimation of basis set superposition error in biomolecular systems, Journal of Chemical Physics 135, 144110.
- Faver, J., Zheng, Z, Merz, K. M. (2012) Statistics-based Model for Basis Set Superposition Error Correction in Large Biomolecules Physical Chemistry Chemical Physics 14, 7795-7799.